An interactive metabolic map of bio-based chemicals.
Trends Biotechnol. 2022; 41: 10-14
The control of flux.
Symp. Soc. Exp. Biol. 1973; 27: 65-104
MPS: an algorithm and data base for metabolic pathway synthesis.
Biotechnol. Lett. 1986; 8: 837-842
Ethanol production by Zymomonas mobilis.
in: Microbial Reactions. Springer, 1982: 37-84
Acetone-butanol fermentation revisited.
Microbiol. Rev. 1986; 50: 484-524
Polymer synthesis by microorganisms: technology and economics.
Trends Biotechnol. 1987; 5: 246-250
Ingram, L.O. et al. Ethanol production by Escherichia coli strains co-expressing Zymomonas PDC and ADH genes, US5000000A, University of Florida Research Foundation Inc
Cloning of the Alcaligenes eutrophus genes for synthesis of poly-beta-hydroxybutyric acid (PHB) and synthesis of PHB in Escherichia coli.
J. Bacteriol. 1988; 170: 5837-5847
1,3-Propanediol production by Escherichia coli expressing genes from the Klebsiella pneumoniae dha regulon.
Appl. Environ. Microbiol. 1991; 57: 3541-3546
Production of 2-keto-L-gulonate, an intermediate in L-ascorbate synthesis, by a genetically modified Erwinia herbicola.
Science. 1985; 230: 144-149
Metabolic engineering advances and prospects for amino acid production.
Metab. Eng. 2020; 58: 17-34
Advances in metabolic engineering of Corynebacterium glutamicum to produce high-value active ingredients for food, feed, human health, and well-being.
Essays Biochem. 2021; 65: 197-212
Metabolic Engineering: Principles and Methodologies.
Academic Press, 1998
Construction of plasmids, estimation of plasmid stability, and use of stable plasmids for the production of poly(3-hydroxybutyric acid) by recombinant Escherichia coli.
J. Biotechnol. 1994; 32: 203-211
Production of poly(3-hydroxybutyrate) by fed-batch culture of filamentation-suppressed recombinant Escherichia coli.
Appl. Environ. Microbiol. 1997; 63: 4765-4769
Genetic construction of lactose-utilizing strains of Pseudomonas aeruginosa and their application in biosurfactant production.
Nat. Biotechnol. 1988; 6: 1335-1339
Efficient degradation of trichloroethylene by a recombinant Escherichia coli.
Nat. Biotechnol. 1989; 7: 282-285
Toward a science of metabolic engineering.
Science. 1991; 252: 1668-1675
Whole-genome random sequencing and assembly of Haemophilus influenzae Rd.
Science. 1995; 269: 496-512
Life with 6000 genes.
Science. 1996; 274: 546-567
The complete genome sequence of Escherichia coli K-12.
Science. 1997; 277: 1453-1462
Systems properties of the Haemophilus influenzae Rd metabolic genotype.
J. Biol. Chem. 1999; 274: 17410-17416
The Escherichia coli MG1655 in silico metabolic genotype: its definition, characteristics, and capabilities.
Proc. Natl. Acad. Sci. U. S. A. 2000; 97: 5528-5533
Proteome profiling and its use in metabolic and cellular engineering.
Proteomics. 2003; 3: 2317-2324
Impact of ‘ome’ analyses on inverse metabolic engineering.
Metab. Eng. 2004; 6: 204-211
Global physiological understanding and metabolic engineering of microorganisms based on omics studies.
Appl. Microbiol. Biotechnol. 2005; 68: 567-579
Native-sized recombinant spider silk protein produced in metabolically engineered Escherichia coli results in a strong fiber.
Proc. Natl. Acad. Sci. U. S. A. 2010; 107: 14059-14063
Metabolic engineering of Escherichia coli for enhanced production of succinic acid, based on genome comparison and in silico gene knockout simulation.
Appl. Environ. Microbiol. 2005; 71: 7880-7887
In silico design and adaptive evolution of Escherichia coli for production of lactic acid.
Biotechnol. Bioeng. 2005; 91: 643-648
Identifying gene targets for the metabolic engineering of lycopene biosynthesis in Escherichia coli.
Metab. Eng. 2005; 7: 155-164
In silico identification of gene amplification targets for improvement of lycopene production.
Appl. Environ. Microbiol. 2010; 76: 3097-3105
Metabolic engineering of Escherichia coli for the production of L-valine based on transcriptome analysis and in silico gene knockout simulation.
Proc. Natl. Acad. Sci. U. S. A. 2007; 104: 7797-7802
From zero to hero – design-based systems metabolic engineering of Corynebacterium glutamicum for L-lysine production.
Metab. Eng. 2011; 13: 159-168
Improving the phenotype predictions of a yeast genome-scale metabolic model by incorporating enzymatic constraints.
Mol. Syst. Biol. 2017; 13: 935
Genome-scale models of metabolism and gene expression extend and refine growth phenotype prediction.
Mol. Syst. Biol. 2013; 9: 693
Current status and applications of genome-scale metabolic models.
Genome Biol. 2019; 20: 121
Reconstructing organisms in silico: genome-scale models and their emerging applications.
Nat. Rev. Microbiol. 2020; 18: 731-743
A TALE nuclease architecture for efficient genome editing.
Nat. Biotechnol. 2011; 29: 143-148
Genome engineering with zinc-finger nucleases.
Genetics. 2011; 188: 773-782
Programming cells by multiplex genome engineering and accelerated evolution.
Nature. 2009; 460: 894-898
RNA-guided editing of bacterial genomes using CRISPR-Cas systems.
Nat. Biotechnol. 2013; 31: 233-239
Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs.
Nat. Biotechnol. 2013; 31: 170-174
Gene expression knockdown by modulating synthetic small RNA expression in Escherichia coli.
Cell Syst. 2017; 5: 418-426
Synthetic CRISPR-Cas gene activators for transcriptional reprogramming in bacteria.
Nat. Commun. 2018; 9: 2489
Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression.
Cell. 2021; 184: 844
Promoter engineering: recent advances in controlling transcription at the most fundamental level.
Biotechnol. J. 2013; 8: 46-58
Precise and reliable gene expression via standard transcription and translation initiation elements.
Nat. Methods. 2013; 10: 354-360
Automated design of synthetic ribosome binding sites to control protein expression.
Nat. Biotechnol. 2009; 27: 946-950
Isoprenoid pathway optimization for Taxol precursor overproduction in Escherichia coli.
Science. 2010; 330: 70-74
Synthetic protein scaffolds provide modular control over metabolic flux.
Nat. Biotechnol. 2009; 27: 753-759
Compartmentalization of metabolic pathways in yeast mitochondria improves the production of branched-chain alcohols.
Nat. Biotechnol. 2013; 31: 335-341
Solution structure of a bacterial microcompartment targeting peptide and its application in the construction of an ethanol bioreactor.
ACS Synth. Biol. 2014; 3: 454-465
Systems metabolic engineering of microorganisms for natural and non-natural chemicals.
Nat. Chem. Biol. 2012; 8: 536-546
Systems strategies for developing industrial microbial strains.
Nat. Biotechnol. 2015; 33: 1061-1072
A comprehensive metabolic map for production of bio-based chemicals.
Nat. Catal. 2019; 2: 18-33
Retrosynthetic design of metabolic pathways to chemicals not found in nature.
Curr. Opin. Syst. Biol. 2019; 14: 82-107
Metabolic engineering of Escherichia coli for direct production of 1,4-butanediol.
Nat. Chem. Biol. 2011; 7: 445-452
Expanding ester biosynthesis in Escherichia coli.
Nat. Chem. Biol. 2014; 10: 259-265
Metabolic engineering of Escherichia coli for the production of four-, five- and six-carbon lactams.
Metab. Eng. 2017; 41: 82-91
One-step fermentative production of aromatic polyesters from glucose by metabolically engineered Escherichia coli strains.
Nat. Commun. 2018; 9: 79
Combinatorial and computational challenges for biocatalyst design.
Nature. 2001; 409: 253-257
Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels.
Nature. 2008; 451: 86-89
Metabolic engineering of Escherichia coli for the production of polylactic acid and its copolymers.
Biotechnol. Bioeng. 2010; 105: 161-171
Systems biology based metabolic engineering for non-natural chemicals.
Biotechnol. Adv. 2019; 37107379
Designing microbial cell factories for the production of chemicals.
JACS Au. 2022; 2: 1781-1799
Production of the antimalarial drug precursor artemisinic acid in engineered yeast.
Nature. 2006; 440: 940-943
A bacterial platform for fermentative production of plant alkaloids.
Nat. Commun. 2011; 2: 326
Metabolic engineering of Escherichia coli for natural product biosynthesis.
Trends Biotechnol. 2020; 38: 745-765
Machine learning applications in systems metabolic engineering.
Curr. Opin. Biotechnol. 2020; 64: 1-9
Enhanced succinic acid production by Mannheimia employing optimal malate dehydrogenase.
Nat. Commun. 2020; 11: 1970
Glutaric acid production by systems metabolic engineering of an L-lysine-overproducing Corynebacterium glutamicum.
Proc. Natl. Acad. Sci. U. S. A. 2020; 117: 30328-30334
Metabolic engineering of Escherichia coli for producing adipic acid through the reverse adipate-degradation pathway.
Metab. Eng. 2018; 47: 254-262
Molecular characterization of the 1,3-propanediol (1,3-PD) operon of Clostridium butyricum.
Proc. Natl. Acad. Sci. U. S. A. 2003; 100: 5010-5015
Co-fermentation of glycerol and glucose by a co-culture system of engineered Escherichia coli strains for 1,3-propanediol production without vitamin B12 supplementation.
Bioresour. Technol. 2021; 319124218
Construction of a synthetic pathway for the production of 1,3-propanediol from glucose.
Sci. Rep. 2019; 9: 11576
Metabolic engineering of a homoserine-derived non-natural pathway for the de novo production of 1,3-propanediol from glucose.
ACS Synth. Biol. 2019; 8: 587-595
Efficient production of 1,3-propanediol from diverse carbohydrates via a non-natural pathway using 3-hydroxypropionic acid as an intermediate.
ACS Synth. Biol. 2021; 10: 478-486
Systems metabolic engineering of Corynebacterium glutamicum for high-level production of 1,3-propanediol from glucose and xylose.
Metab. Eng. 2022; 70: 79-88
Rational flux-tuning of Halomonas bluephagenesis for co-production of bioplastic PHB and ectoine.
Nat. Commun. 2020; 11: 3313
Halomonas as a chassis.
Essays Biochem. 2021; 65: 393-403
Biosynthesis of polyhydroxyalkanoates containing 2-hydroxy-4-methylvalerate and 2-hydroxy-3-phenylpropionate units from a related or unrelated carbon source.
J. Biosci. Bioeng. 2018; 125: 295-300
Biosynthesis of poly(2-hydroxyisovalerate-co-lactate) by metabolically engineered Escherichia coli.
Biotechnol. J. 2016; 11: 1572-1585
One-step fermentative production of poly(lactate-co-glycolate) from carbohydrates in Escherichia coli.
Nat. Biotechnol. 2016; 34: 435-440
Metabolic engineering for the synthesis of polyesters: a 100-year journey from polyhydroxyalkanoates to non-natural microbial polyesters.
Metab. Eng. 2020; 58: 47-81
Biosynthesizing structurally diverse diols via a general route combining oxidative and reductive formations of OH-groups.
Nat. Commun. 2022; 13: 1595
Microbial production of multiple short-chain primary amines via retrobiosynthesis.
Nat. Commun. 2021; 12: 173
The role of microbes in biofuel production.
in: Kumar N. Biofuel from Microbes and Plants: Green Energy Alternative. 1st edn. CRC Press, 2021: 63
Microbial biotechnological approaches: renewable bioprocessing for the future energy systems.
Microb. Cell Factories. 2021; 20: 55
Biofuels for a sustainable future.
Cell. 2021; 184: 1636-1647
Microbial production of advanced biofuels.
Nat. Rev. Microbiol. 2021; 19: 701-715
Bioethanol.
Curr. Opin. Chem. Biol. 2006; 10: 141-146
One hundred years of clostridial butanol fermentation.
FEMS Microbiol. Lett. 2016; 363e02683–18
Bioethanol production from renewable sources: current perspectives and technological progress.
Renew. Sustain. Energy Rev. 2017; 71: 475-501
Recent advances and state-of-the-art strategies in strain and process engineering for biobutanol production by Clostridium acetobutylicum.
Biotechnol. Adv. 2017; 35: 310-322
Enhanced butanol production obtained by reinforcing the direct butanol-forming route in Clostridium acetobutylicum.
mBio. 2012; 3e00314–12
Genetic engineering of ethanol production in Escherichia coli.
Appl. Environ. Microbiol. 1987; 53: 2420-2425
Genetic improvement of Escherichia coli for ethanol production: chromosomal integration of Zymomonas mobilis genes encoding pyruvate decarboxylase and alcohol dehydrogenase II.
Appl. Environ. Microbiol. 1991; 57: 893-900
Expression of different levels of enzymes from the Pichia stipitis XYL1 and XYL2 genes in Saccharomyces cerevisiae and its effects on product formation during xylose utilisation.
Appl. Microbiol. Biotechnol. 1997; 48: 218-224
In silico aided metabolic engineering of Saccharomyces cerevisiae for improved bioethanol production.
Metab. Eng. 2006; 8: 102-111
Xylose isomerase overexpression along with engineering of the pentose phosphate pathway and evolutionary engineering enable rapid xylose utilization and ethanol production by Saccharomyces cerevisiae.
Metab. Eng. 2012; 14: 611-622
Elimination of glycerol production in anaerobic cultures of a Saccharomyces cerevisiae strain engineered to use acetic acid as an electron acceptor.
Appl. Environ. Microbiol. 2010; 76: 190-195
Engineering yeast transcription machinery for improved ethanol tolerance and production.
Science. 2006; 314: 1565-1568
Enhanced biofuel production through coupled acetic acid and xylose consumption by engineered yeast.
Nat. Commun. 2013; 4: 2580
Microbial production of short-chain alkanes.
Nature. 2013; 502: 571-574
Modular and selective biosynthesis of gasoline-range alkanes.
Metab. Eng. 2016; 33: 28-40
Engineered reversal of the beta-oxidation cycle for the synthesis of fuels and chemicals.
Nature. 2011; 476: 355-359
Modular optimization of multi-gene pathways for fatty acids production in E. coli.
Nat. Commun. 2013; 4: 1409
Production of fatty acid-derived oleochemicals and biofuels by synthetic yeast cell factories.
Nat. Commun. 2016; 7: 11709
Reprogramming yeast metabolism from alcoholic fermentation to lipogenesis.
Cell. 2018; 174: 1549-1558
Metabolic reconfiguration enables synthetic reductive metabolism in yeast.
Nat. Metab. 2022; 4: 1551-1559
Engineering of an oleaginous bacterium for the production of fatty acids and fuels.
Nat. Chem. Biol. 2019; 15: 721-729
Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids.
Nat. Biotechnol. 2012; 30: 354-359
Microbial production of fatty-acid-derived fuels and chemicals from plant biomass.
Nature. 2010; 463: 559-562
Engineering Yarrowia lipolytica as a platform for synthesis of drop-in transportation fuels and oleochemicals.
Proc. Natl. Acad. Sci. U. S. A. 2016; 113: 10848-10853
Synthesis of FAEEs from glycerol in engineered Saccharomyces cerevisiae using endogenously produced ethanol by heterologous expression of an unspecific bacterial acyltransferase.
Biotechnol. Bioeng. 2012; 109: 110-115
Carotenoid-based phenotypic screen of the yeast deletion collection reveals new genes with roles in isoprenoid production.
Metab. Eng. 2013; 15: 174-183
Rewriting yeast central carbon metabolism for industrial isoprenoid production.
Nature. 2016; 537: 694-697
Transforming yeast peroxisomes into microfactories for the efficient production of high-value isoprenoids.
Proc. Natl. Acad. Sci. U. S. A. 2020; 117: 31789-31799
Peroxisome compartmentalization of a toxic enzyme improves alkaloid production.
Nat. Chem. Biol. 2021; 17: 96-103
Engineering an iterative polyketide pathway in Escherichia coli results in single-form alkene and alkane overproduction.
Metab. Eng. 2015; 28: 82-90
Biosynthesis of polycyclopropanated high energy biofuels.
Joule. 2022; 6: 1590-1605
Drop-in biofuel production using fatty acid photodecarboxylase from Chlorella variabilis in the oleaginous yeast Yarrowia lipolytica.
Biotechnol. Biofuels. 2019; 12: 202
An algal photoenzyme converts fatty acids to hydrocarbons.
Science. 2017; 357: 903-907
Synthesis of high-titer alka(e)nes in Yarrowia lipolytica is enabled by a discovered mechanism.
Nat. Commun. 2020; 11: 6198
Lipid production in Yarrowia lipolytica is maximized by engineering cytosolic redox metabolism.
Nat. Biotechnol. 2017; 35: 173-177
Microbial production and consumption of hydrocarbons in the global ocean.
Nat. Microbiol. 2021; 6: 489-498
Synthetic biology: a new frontier in food production.
Trends Biotechnol. 2022; 40: 781-803
Microbial production of methyl anthranilate, a grape flavor compound.
Proc. Natl. Acad. Sci. U. S. A. 2019; 116: 10749-10756
Industrial brewing yeast engineered for the production of primary flavor determinants in hopped beer.
Nat. Commun. 2018; 9: 965
Production of rainbow colorants by metabolically engineered Escherichia coli.
Adv. Sci. 2021; 8e2100743
Production of natural colorants by metabolically engineered microorganisms.
Trends Chem. 2022; 4: 608-626
Production of carminic acid by metabolically engineered Escherichia coli.
J. Am. Chem. Soc. 2021; 143: 5364-5377
Metabolic engineering of Escherichia coli for secretory production of free haem.
Nat. Catal. 2018; 1: 720-728
Genome-scale modeling drives 70-fold improvement of intracellular heme production in Saccharomyces cerevisiae.
Proc. Natl. Acad. Sci. U. S. A. 2022; 119e2108245119
Improved production of heme using metabolically engineered Escherichia coli.
Biotechnol. Bioeng. 2022; 119: 3178-3193
antiSMASH 6.0: improving cluster detection and comparison capabilities.
Nucleic Acids Res. 2021; 49: W29-W35
Comprehensive prediction of secondary metabolite structure and biological activity from microbial genome sequences.
Nat. Commun. 2020; 11: 6058
Deep learning driven biosynthetic pathways navigation for natural products with BioNavi-NP.
Nat. Commun. 2022; 13: 3342
A computational workflow for the expansion of heterologous biosynthetic pathways to natural product derivatives.
Nat. Commun. 2021; 12: 1760
Metabolic engineering of Escherichia coli with electron channelling for the production of natural products.
Nat. Catal. 2022; : 1-12
Complete biosynthesis of opioids in yeast.
Science. 2015; 349: 1095-1100
Complete biosynthesis of cannabinoids and their unnatural analogues in yeast.
Nature. 2019; 567: 123-126
Biosynthesis of medicinal tropane alkaloids in yeast.
Nature. 2020; 585: 614-619
Targeting pathway expression to subcellular organelles improves astaxanthin synthesis in Yarrowia lipolytica.
Metab. Eng. 2021; 68: 152-161
Removal of lycopene substrate inhibition enables high carotenoid productivity in Yarrowia lipolytica.
Nat. Commun. 2022; 13: 572
A microbial supply chain for production of the anti-cancer drug vinblastine.
Nature. 2022; 609: 341-347
Current perspective of innovative strategies for bioremediation of organic pollutants from wastewater.
Bioresour. Technol. 2022; 344126305
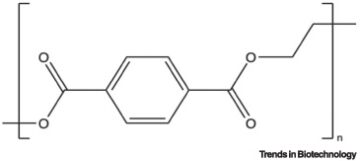
Techno-economic, life-cycle, and socioeconomic impact analysis of enzymatic recycling of poly(ethylene terephthalate).
Joule. 2021; 5: 2479-2503
A bacterium that degrades and assimilates poly(ethylene terephthalate).
Science. 2016; 351: 1196-1199
Characterization and engineering of a plastic-degrading aromatic polyesterase.
Proc. Natl. Acad. Sci. U. S. A. 2018; 115: E4350-E4357
Structural insight into molecular mechanism of poly(ethylene terephthalate) degradation.
Nat. Commun. 2018; 9: 382
Rational protein engineering of thermo-stable PETase from Ideonella sakaiensis for highly efficient PET degradation.
ACS Catal. 2019; 9: 3519-3526
Structural bioinformatics-based protein engineering of thermo-stable PETase from Ideonella sakaiensis.
Enzym. Microb. Technol. 2020; 141109656
Computational redesign of a PETase for plastic biodegradation under ambient condition by the GRAPE strategy.
ACS Catal. 2021; 11: 1340-1350
Structure of the plastic-degrading Ideonella sakaiensis MHETase bound to a substrate.
Nat. Commun. 2019; 10: 1717
Decomposition of the PET film by MHETase using Exo-PETase function.
ACS Catal. 2020; 10: 4805-4812
Characterization and engineering of a two-enzyme system for plastics depolymerization.
Proc. Natl. Acad. Sci. U. S. A. 2020; 117: 25476-25485
Directed evolution of an efficient and thermostable PET depolymerase.
Nat. Catal. 2022; 5: 673-681
Machine learning-aided engineering of hydrolases for PET depolymerization.
Nature. 2022; 604: 662-667
An engineered PET depolymerase to break down and recycle plastic bottles.
Nature. 2020; 580: 216-219
Enzyme discovery and engineering for sustainable plastic recycling.
Trends Biotechnol. 2022; 40: 22-37
Polyvinyl chloride degradation by a bacterium isolated from the gut of insect larvae.
Nat. Commun. 2022; 13: 5360
Wax worm saliva and the enzymes therein are the key to polyethylene degradation by Galleria mellonella.
Nat. Commun. 2022; 13: 5568
Towards bio-upcycling of polyethylene terephthalate.
Metab. Eng. 2021; 66: 167-178
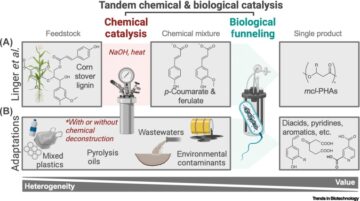
Mixed plastics waste valorization through tandem chemical oxidation and biological funneling.
Science. 2022; 378: 207-211
New applications of synthetic biology tools for cyanobacterial metabolic engineering.
Front. Bioeng. Biotechnol. 2019; 7: 33
Advances in systems metabolic engineering of autotrophic carbon oxide-fixing biocatalysts towards a circular economy.
Metab. Eng. 2022; 71: 117-141
Direct photosynthetic recycling of carbon dioxide to isobutyraldehyde.
Nat. Biotechnol. 2009; 27: 1177-1180
Carbon-negative production of acetone and isopropanol by gas fermentation at industrial pilot scale.
Nat. Biotechnol. 2022; 40: 335-344
Third-generation biorefineries as the means to produce fuels and chemicals from CO2.
Nat. Catal. 2020; 3: 274-288
Conversion of Escherichia coli to generate all biomass carbon from CO2.
Cell. 2019; 179: 1255-1263 e12
Escherichia coli is engineered to grow on CO2 and formic acid.
Nat. Microbiol. 2020; 5: 1459-1463
Converting Escherichia coli to a synthetic methylotroph growing solely on methanol.
Cell. 2020; 182: 933-946
Integrated electromicrobial conversion of CO2 to higher alcohols.
Science. 2012; 335: 1596
From CO2 to bioplastic – coupling the electrochemical CO2 reduction with a microbial product generation by drop-in electrolysis.
ChemSusChem. 2020; 13: 4086-4093
Nonchromosomal antibiotic resistance in bacteria: genetic transformation of Escherichia coli by R-factor DNA.
Proc. Natl. Acad. Sci. U. S. A. 1972; 69: 2110-2114
Construction of biologically functional bacterial plasmids in vitro.
Proc. Natl. Acad. Sci. U. S. A. 1973; 70: 3240-3244
DNA sequencing with chain-terminating inhibitors.
Proc. Natl. Acad. Sci. U. S. A. 1977; 74: 5463-5467
Expression in Escherichia coli of a chemically synthesized gene for the hormone somatostatin.
Science. 1977; 198: 1056-1063
Expression in Escherichia coli of chemically synthesized genes for human insulin.
Proc. Natl. Acad. Sci. U. S. A. 1979; 76: 106-110
PCR: thirty-five years and counting.
Science. 2018; 360: 670-672
Basic local alignment search tool.
J. Mol. Biol. 1990; 215: 403-410
Network analysis of intermediary metabolism using linear optimization. I. Development of mathematical formalism.
J. Theor. Biol. 1992; 154: 421-454
A simple and efficient method for direct gene deletion in Saccharomyces cerevisiae.
Nucleic Acids Res. 1993; 21: 3329-3330
On elementary flux modes in biochemical reaction systems at steady state.
J. Biol. Syst. 2011; 02: 165-182
Intracellular flux analysis in hybridomas using mass balances and in vitro 13C NMR.
Biotechnol. Bioeng. 1995; 45: 292-303
A new logic for DNA engineering using recombination in Escherichia coli.
Nat. Genet. 1998; 20: 123-128
One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products.
Proc. Natl. Acad. Sci. U. S. A. 2000; 97: 6640-6645
Genome shuffling leads to rapid phenotypic improvement in bacteria.
Nature. 2002; 415: 644-646
Genome shuffling of Lactobacillus for improved acid tolerance.
Nat. Biotechnol. 2002; 20: 707-712
Analysis of optimality in natural and perturbed metabolic networks.
Proc. Natl. Acad. Sci. U. S. A. 2002; 99: 15112-15117
The effects of alternate optimal solutions in constraint-based genome-scale metabolic models.
Metab. Eng. 2003; 5: 264-276
OptKnock: a bilevel programming framework for identifying gene knockout strategies for microbial strain optimization.
Biotechnol. Bioeng. 2003; 84: 647-657
Total biosynthesis of hydrocortisone from a simple carbon source in yeast.
Nat. Biotechnol. 2003; 21: 143-149
Genome sequencing in microfabricated high-density picolitre reactors.
Nature. 2005; 437: 376-380
Complete chemical synthesis, assembly, and cloning of a Mycoplasma genitalium genome.
Science. 2008; 319: 1215-1220
Enzymatic assembly of DNA molecules up to several hundred kilobases.
Nat. Methods. 2009; 6: 343-345
Three-dimensional structural view of the central metabolic network of Thermotoga maritima.
Science. 2009; 325: 1544-1549
Creation of a bacterial cell controlled by a chemically synthesized genome.
Science. 2010; 329: 52-56
antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences.
Nucleic Acids Res. 2011; 39: W339-W346
A whole-cell computational model predicts phenotype from genotype.
Cell. 2012; 150: 389-401
Synthetic non-oxidative glycolysis enables complete carbon conservation.
Nature. 2013; 502: 693-697
Computational protein design enables a novel one-carbon assimilation pathway.
Proc. Natl. Acad. Sci. U. S. A. 2015; 112: 3704-3709
Genomes to natural products PRediction Informatics for Secondary Metabolomes (PRISM).
Nucleic Acids Res. 2015; 43: 9645-9662
A genome-scale Escherichia coli kinetic metabolic model k-ecoli457 satisfying flux data for multiple mutant strains.
Nat. Commun. 2016; 7: 13806
Genetic circuit design automation.
Science. 2016; 352aac7341
Assimilation of formic acid and CO2 by engineered Escherichia coli equipped with reconstructed one-carbon assimilation pathways.
Proc. Natl. Acad. Sci. U. S. A. 2018; 115: E9271-E9279
Improved protein structure prediction using potentials from deep learning.
Nature. 2020; 577: 706-710
Deep learning enables high-quality and high-throughput prediction of enzyme commission numbers.
Proc. Natl. Acad. Sci. U. S. A. 2019; 116: 13996-14001
Simultaneous cross-evaluation of heterogeneous E. coli datasets via mechanistic simulation.
Science. 2020; 369eaav3751
Highly accurate protein structure prediction with AlphaFold.
Nature. 2021; 596: 583-589
Accurate prediction of protein structures and interactions using a three-track neural network.
Science. 2021; 373: 871-876
- SEO Powered Content & PR Distribution. Get Amplified Today.
- Platoblockchain. Web3 Metaverse Intelligence. Knowledge Amplified. Access Here.
- Source: https://www.cell.com/trends/biotechnology/fulltext/S0167-7799(22)00342-0?rss=yes
- 1
- a
- accelerated
- accurate
- active
- activity
- advanced
- advances
- Alcohol
- algorithm
- All
- alternative
- Ambient
- analysis
- and
- Application
- applications
- approaches
- architecture
- Assembly
- authors
- Automation
- Bacteria
- balances
- base
- based
- beer
- binding
- biology
- biomass
- bound
- Break
- capabilities
- carbon
- carbon dioxide
- Cells
- central
- chain
- challenges
- characteristics
- chassis
- chemical
- chemicals
- circular economy
- Cluster
- collection
- combining
- commission
- comparison
- complete
- Compound
- comprehensive
- condition
- CONSERVATION
- constraints
- construction
- consumption
- control
- controlled
- controlling
- Conversion
- coupled
- CRISPR
- Culture
- Current
- data
- datasets
- deep
- deep learning
- Derivatives
- Derived
- Design
- Detection
- developing
- Development
- different
- direct
- discovered
- discovery
- diverse
- dna
- down
- driven
- drug
- during
- dynamic
- Economics
- economy
- effects
- efficient
- elements
- emerging
- enable
- enabled
- enables
- energy
- Engineering
- enhanced
- equipped
- evolution
- expansion
- extend
- factories
- Film
- florida
- FLUX
- food
- formation
- found
- Foundation
- Framework
- Free
- from
- Frontier
- function
- functional
- fundamental
- future
- GAS
- General
- generate
- generation
- genome
- Global
- Green
- green energy
- Grow
- Growing
- Growth
- Health
- Hero
- High
- high-level
- high-quality
- higher
- highly
- HTTPS
- human
- Identification
- identifying
- Impact
- improved
- improvement
- improves
- improving
- in
- incorporating
- industrial
- innovative
- insight
- integration
- interactions
- interactive
- Intermediate
- isolated
- journey
- Key
- Leads
- learning
- Level
- levels
- life-cycle
- List
- local
- machinery
- map
- Mass
- mathematical
- means
- mechanism
- medicinal
- Methanol
- method
- methodologies
- methods
- Mitochondria
- model
- modeling
- models
- modified
- modular
- MOL
- molecular
- most
- multiple
- Natural
- Nature
- Navigation
- network
- networks
- Neural
- neural network
- New
- novel
- numbers
- obtained
- ocean
- optimal
- optimization
- organic
- PCR
- perspective
- perspectives
- phenotype
- pilot
- plants
- plastic
- plastics
- platform
- plato
- Plato Data Intelligence
- PlatoData
- precursor
- prediction
- Predictions
- Predicts
- primary
- principles
- process
- produce
- Produced
- Product
- Production
- productivity
- Products
- profiling
- Programming
- Progress
- properties
- prospects
- Protein
- provide
- random
- rapid
- reaction
- reactions
- recent
- recycling
- redesign
- regulatory
- related
- reliable
- Renewable
- research
- Resistance
- Results
- Reveals
- Reversal
- reverse
- RNA
- Role
- roles
- Route
- Saliva
- Scale
- SCI
- Science
- Screen
- Search
- secondary
- selective
- Sequence
- several
- silk
- Simple
- simulation
- Sites
- small
- socioeconomic
- Solutions
- Source
- Sources
- Spider Silk
- Stability
- stable
- standard
- State
- state-of-the-art
- Status
- steady
- Strains
- strategies
- Strategy
- strong
- structural
- structure
- studies
- supply
- supply chain
- Sustainability
- sustainable
- synthetic
- system
- Systems
- Tandem
- targeting
- targets
- technological
- Technology
- The
- The Future
- their
- therein
- Through
- to
- tolerance
- tool
- tools
- towards
- Transformation
- Translation
- transportation
- under
- understanding
- units
- university
- use
- via
- View
- W
- Waste
- without
- workflow
- worm
- X
- years
- zephyrnet
- zero